More python tips#
# import the pandas package
import pandas as pd
# import numpy
import numpy as np
# import the package we'll use for plotting
import matplotlib.pyplot as plt
# this tells the jupyter notebook to show plots "inline" with other output here in the notebook
%matplotlib inline
Opening data files (with pandas):#
# Location of the data file
Skykomish_data_file = 'Skykomish_peak_flow_12134500_skykomish_river_near_gold_bar.xlsx'
# Use pandas.read_excel() function to open this file.
Skykomish_data = pd.read_excel(Skykomish_data_file)
/opt/hostedtoolcache/Python/3.7.17/x64/lib/python3.7/site-packages/openpyxl/worksheet/_reader.py:329: UserWarning: Unknown extension is not supported and will be removed
warn(msg)
# Now we can see the dataset we loaded:
Skykomish_data
| date of peak | water year | peak value (cfs) | gage_ht (feet) | |
|---|---|---|---|---|
| 0 | 1928-10-09 | 1929 | 18800 | 10.55 |
| 1 | 1930-02-05 | 1930 | 15800 | 10.44 |
| 2 | 1931-01-28 | 1931 | 35100 | 14.08 |
| 3 | 1932-02-26 | 1932 | 83300 | 20.70 |
| 4 | 1932-11-13 | 1933 | 72500 | 19.50 |
| ... | ... | ... | ... | ... |
| 86 | 2015-11-17 | 2016 | 95900 | 21.73 |
| 87 | 2016-10-20 | 2017 | 41000 | 15.76 |
| 88 | 2017-11-23 | 2018 | 54200 | 17.50 |
| 89 | 2018-11-02 | 2019 | 35200 | 14.98 |
| 90 | 2020-02-01 | 2020 | 72200 | 19.50 |
91 rows × 4 columns
# Look at a single column, using the header name
Skykomish_data['water year']
0 1929
1 1930
2 1931
3 1932
4 1933
...
86 2016
87 2017
88 2018
89 2019
90 2020
Name: water year, Length: 91, dtype: int64
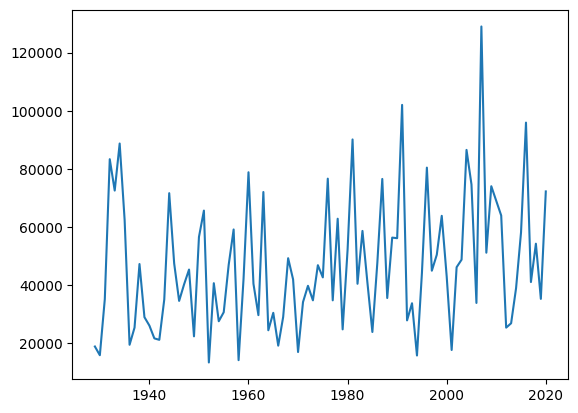
# Plot data, specifying our x and y with column header names
plt.plot(Skykomish_data['water year'],Skykomish_data['peak value (cfs)'])
[<matplotlib.lines.Line2D at 0x7f51cfb3c510>]
How to find help/documentation?#
help(np.mean)
Help on function mean in module numpy:
mean(a, axis=None, dtype=None, out=None, keepdims=<no value>, *, where=<no value>)
Compute the arithmetic mean along the specified axis.
Returns the average of the array elements. The average is taken over
the flattened array by default, otherwise over the specified axis.
`float64` intermediate and return values are used for integer inputs.
Parameters
----------
a : array_like
Array containing numbers whose mean is desired. If `a` is not an
array, a conversion is attempted.
axis : None or int or tuple of ints, optional
Axis or axes along which the means are computed. The default is to
compute the mean of the flattened array.
.. versionadded:: 1.7.0
If this is a tuple of ints, a mean is performed over multiple axes,
instead of a single axis or all the axes as before.
dtype : data-type, optional
Type to use in computing the mean. For integer inputs, the default
is `float64`; for floating point inputs, it is the same as the
input dtype.
out : ndarray, optional
Alternate output array in which to place the result. The default
is ``None``; if provided, it must have the same shape as the
expected output, but the type will be cast if necessary.
See :ref:`ufuncs-output-type` for more details.
keepdims : bool, optional
If this is set to True, the axes which are reduced are left
in the result as dimensions with size one. With this option,
the result will broadcast correctly against the input array.
If the default value is passed, then `keepdims` will not be
passed through to the `mean` method of sub-classes of
`ndarray`, however any non-default value will be. If the
sub-class' method does not implement `keepdims` any
exceptions will be raised.
where : array_like of bool, optional
Elements to include in the mean. See `~numpy.ufunc.reduce` for details.
.. versionadded:: 1.20.0
Returns
-------
m : ndarray, see dtype parameter above
If `out=None`, returns a new array containing the mean values,
otherwise a reference to the output array is returned.
See Also
--------
average : Weighted average
std, var, nanmean, nanstd, nanvar
Notes
-----
The arithmetic mean is the sum of the elements along the axis divided
by the number of elements.
Note that for floating-point input, the mean is computed using the
same precision the input has. Depending on the input data, this can
cause the results to be inaccurate, especially for `float32` (see
example below). Specifying a higher-precision accumulator using the
`dtype` keyword can alleviate this issue.
By default, `float16` results are computed using `float32` intermediates
for extra precision.
Examples
--------
>>> a = np.array([[1, 2], [3, 4]])
>>> np.mean(a)
2.5
>>> np.mean(a, axis=0)
array([2., 3.])
>>> np.mean(a, axis=1)
array([1.5, 3.5])
In single precision, `mean` can be inaccurate:
>>> a = np.zeros((2, 512*512), dtype=np.float32)
>>> a[0, :] = 1.0
>>> a[1, :] = 0.1
>>> np.mean(a)
0.54999924
Computing the mean in float64 is more accurate:
>>> np.mean(a, dtype=np.float64)
0.55000000074505806 # may vary
Specifying a where argument:
>>> a = np.array([[5, 9, 13], [14, 10, 12], [11, 15, 19]])
>>> np.mean(a)
12.0
>>> np.mean(a, where=[[True], [False], [False]])
9.0
np.mean?
help(pd.read_excel)
Help on function read_excel in module pandas.io.excel._base:
read_excel(io, sheet_name=0, header=0, names=None, index_col=None, usecols=None, squeeze=False, dtype: 'DtypeArg | None' = None, engine=None, converters=None, true_values=None, false_values=None, skiprows=None, nrows=None, na_values=None, keep_default_na=True, na_filter=True, verbose=False, parse_dates=False, date_parser=None, thousands=None, comment=None, skipfooter=0, convert_float=None, mangle_dupe_cols=True, storage_options: 'StorageOptions' = None)
Read an Excel file into a pandas DataFrame.
Supports `xls`, `xlsx`, `xlsm`, `xlsb`, `odf`, `ods` and `odt` file extensions
read from a local filesystem or URL. Supports an option to read
a single sheet or a list of sheets.
Parameters
----------
io : str, bytes, ExcelFile, xlrd.Book, path object, or file-like object
Any valid string path is acceptable. The string could be a URL. Valid
URL schemes include http, ftp, s3, and file. For file URLs, a host is
expected. A local file could be: ``file://localhost/path/to/table.xlsx``.
If you want to pass in a path object, pandas accepts any ``os.PathLike``.
By file-like object, we refer to objects with a ``read()`` method,
such as a file handle (e.g. via builtin ``open`` function)
or ``StringIO``.
sheet_name : str, int, list, or None, default 0
Strings are used for sheet names. Integers are used in zero-indexed
sheet positions. Lists of strings/integers are used to request
multiple sheets. Specify None to get all sheets.
Available cases:
* Defaults to ``0``: 1st sheet as a `DataFrame`
* ``1``: 2nd sheet as a `DataFrame`
* ``"Sheet1"``: Load sheet with name "Sheet1"
* ``[0, 1, "Sheet5"]``: Load first, second and sheet named "Sheet5"
as a dict of `DataFrame`
* None: All sheets.
header : int, list of int, default 0
Row (0-indexed) to use for the column labels of the parsed
DataFrame. If a list of integers is passed those row positions will
be combined into a ``MultiIndex``. Use None if there is no header.
names : array-like, default None
List of column names to use. If file contains no header row,
then you should explicitly pass header=None.
index_col : int, list of int, default None
Column (0-indexed) to use as the row labels of the DataFrame.
Pass None if there is no such column. If a list is passed,
those columns will be combined into a ``MultiIndex``. If a
subset of data is selected with ``usecols``, index_col
is based on the subset.
usecols : int, str, list-like, or callable default None
* If None, then parse all columns.
* If str, then indicates comma separated list of Excel column letters
and column ranges (e.g. "A:E" or "A,C,E:F"). Ranges are inclusive of
both sides.
* If list of int, then indicates list of column numbers to be parsed.
* If list of string, then indicates list of column names to be parsed.
* If callable, then evaluate each column name against it and parse the
column if the callable returns ``True``.
Returns a subset of the columns according to behavior above.
squeeze : bool, default False
If the parsed data only contains one column then return a Series.
dtype : Type name or dict of column -> type, default None
Data type for data or columns. E.g. {'a': np.float64, 'b': np.int32}
Use `object` to preserve data as stored in Excel and not interpret dtype.
If converters are specified, they will be applied INSTEAD
of dtype conversion.
engine : str, default None
If io is not a buffer or path, this must be set to identify io.
Supported engines: "xlrd", "openpyxl", "odf", "pyxlsb".
Engine compatibility :
- "xlrd" supports old-style Excel files (.xls).
- "openpyxl" supports newer Excel file formats.
- "odf" supports OpenDocument file formats (.odf, .ods, .odt).
- "pyxlsb" supports Binary Excel files.
.. versionchanged:: 1.2.0
The engine `xlrd <https://xlrd.readthedocs.io/en/latest/>`_
now only supports old-style ``.xls`` files.
When ``engine=None``, the following logic will be
used to determine the engine:
- If ``path_or_buffer`` is an OpenDocument format (.odf, .ods, .odt),
then `odf <https://pypi.org/project/odfpy/>`_ will be used.
- Otherwise if ``path_or_buffer`` is an xls format,
``xlrd`` will be used.
- Otherwise if ``path_or_buffer`` is in xlsb format,
``pyxlsb`` will be used.
.. versionadded:: 1.3.0
- Otherwise ``openpyxl`` will be used.
.. versionchanged:: 1.3.0
converters : dict, default None
Dict of functions for converting values in certain columns. Keys can
either be integers or column labels, values are functions that take one
input argument, the Excel cell content, and return the transformed
content.
true_values : list, default None
Values to consider as True.
false_values : list, default None
Values to consider as False.
skiprows : list-like, int, or callable, optional
Line numbers to skip (0-indexed) or number of lines to skip (int) at the
start of the file. If callable, the callable function will be evaluated
against the row indices, returning True if the row should be skipped and
False otherwise. An example of a valid callable argument would be ``lambda
x: x in [0, 2]``.
nrows : int, default None
Number of rows to parse.
na_values : scalar, str, list-like, or dict, default None
Additional strings to recognize as NA/NaN. If dict passed, specific
per-column NA values. By default the following values are interpreted
as NaN: '', '#N/A', '#N/A N/A', '#NA', '-1.#IND', '-1.#QNAN', '-NaN', '-nan',
'1.#IND', '1.#QNAN', '<NA>', 'N/A', 'NA', 'NULL', 'NaN', 'n/a',
'nan', 'null'.
keep_default_na : bool, default True
Whether or not to include the default NaN values when parsing the data.
Depending on whether `na_values` is passed in, the behavior is as follows:
* If `keep_default_na` is True, and `na_values` are specified, `na_values`
is appended to the default NaN values used for parsing.
* If `keep_default_na` is True, and `na_values` are not specified, only
the default NaN values are used for parsing.
* If `keep_default_na` is False, and `na_values` are specified, only
the NaN values specified `na_values` are used for parsing.
* If `keep_default_na` is False, and `na_values` are not specified, no
strings will be parsed as NaN.
Note that if `na_filter` is passed in as False, the `keep_default_na` and
`na_values` parameters will be ignored.
na_filter : bool, default True
Detect missing value markers (empty strings and the value of na_values). In
data without any NAs, passing na_filter=False can improve the performance
of reading a large file.
verbose : bool, default False
Indicate number of NA values placed in non-numeric columns.
parse_dates : bool, list-like, or dict, default False
The behavior is as follows:
* bool. If True -> try parsing the index.
* list of int or names. e.g. If [1, 2, 3] -> try parsing columns 1, 2, 3
each as a separate date column.
* list of lists. e.g. If [[1, 3]] -> combine columns 1 and 3 and parse as
a single date column.
* dict, e.g. {'foo' : [1, 3]} -> parse columns 1, 3 as date and call
result 'foo'
If a column or index contains an unparsable date, the entire column or
index will be returned unaltered as an object data type. If you don`t want to
parse some cells as date just change their type in Excel to "Text".
For non-standard datetime parsing, use ``pd.to_datetime`` after ``pd.read_excel``.
Note: A fast-path exists for iso8601-formatted dates.
date_parser : function, optional
Function to use for converting a sequence of string columns to an array of
datetime instances. The default uses ``dateutil.parser.parser`` to do the
conversion. Pandas will try to call `date_parser` in three different ways,
advancing to the next if an exception occurs: 1) Pass one or more arrays
(as defined by `parse_dates`) as arguments; 2) concatenate (row-wise) the
string values from the columns defined by `parse_dates` into a single array
and pass that; and 3) call `date_parser` once for each row using one or
more strings (corresponding to the columns defined by `parse_dates`) as
arguments.
thousands : str, default None
Thousands separator for parsing string columns to numeric. Note that
this parameter is only necessary for columns stored as TEXT in Excel,
any numeric columns will automatically be parsed, regardless of display
format.
comment : str, default None
Comments out remainder of line. Pass a character or characters to this
argument to indicate comments in the input file. Any data between the
comment string and the end of the current line is ignored.
skipfooter : int, default 0
Rows at the end to skip (0-indexed).
convert_float : bool, default True
Convert integral floats to int (i.e., 1.0 --> 1). If False, all numeric
data will be read in as floats: Excel stores all numbers as floats
internally.
.. deprecated:: 1.3.0
convert_float will be removed in a future version
mangle_dupe_cols : bool, default True
Duplicate columns will be specified as 'X', 'X.1', ...'X.N', rather than
'X'...'X'. Passing in False will cause data to be overwritten if there
are duplicate names in the columns.
storage_options : dict, optional
Extra options that make sense for a particular storage connection, e.g.
host, port, username, password, etc., if using a URL that will
be parsed by ``fsspec``, e.g., starting "s3://", "gcs://". An error
will be raised if providing this argument with a local path or
a file-like buffer. See the fsspec and backend storage implementation
docs for the set of allowed keys and values.
.. versionadded:: 1.2.0
Returns
-------
DataFrame or dict of DataFrames
DataFrame from the passed in Excel file. See notes in sheet_name
argument for more information on when a dict of DataFrames is returned.
See Also
--------
DataFrame.to_excel : Write DataFrame to an Excel file.
DataFrame.to_csv : Write DataFrame to a comma-separated values (csv) file.
read_csv : Read a comma-separated values (csv) file into DataFrame.
read_fwf : Read a table of fixed-width formatted lines into DataFrame.
Examples
--------
The file can be read using the file name as string or an open file object:
>>> pd.read_excel('tmp.xlsx', index_col=0) # doctest: +SKIP
Name Value
0 string1 1
1 string2 2
2 #Comment 3
>>> pd.read_excel(open('tmp.xlsx', 'rb'),
... sheet_name='Sheet3') # doctest: +SKIP
Unnamed: 0 Name Value
0 0 string1 1
1 1 string2 2
2 2 #Comment 3
Index and header can be specified via the `index_col` and `header` arguments
>>> pd.read_excel('tmp.xlsx', index_col=None, header=None) # doctest: +SKIP
0 1 2
0 NaN Name Value
1 0.0 string1 1
2 1.0 string2 2
3 2.0 #Comment 3
Column types are inferred but can be explicitly specified
>>> pd.read_excel('tmp.xlsx', index_col=0,
... dtype={'Name': str, 'Value': float}) # doctest: +SKIP
Name Value
0 string1 1.0
1 string2 2.0
2 #Comment 3.0
True, False, and NA values, and thousands separators have defaults,
but can be explicitly specified, too. Supply the values you would like
as strings or lists of strings!
>>> pd.read_excel('tmp.xlsx', index_col=0,
... na_values=['string1', 'string2']) # doctest: +SKIP
Name Value
0 NaN 1
1 NaN 2
2 #Comment 3
Comment lines in the excel input file can be skipped using the `comment` kwarg
>>> pd.read_excel('tmp.xlsx', index_col=0, comment='#') # doctest: +SKIP
Name Value
0 string1 1.0
1 string2 2.0
2 None NaN
pd.read_excel?
Writing functions:#
def add_values(x,y):
z = x + y
return z
result = add_values(1,2)
print(result)
3
Import your custom functions:#
#def subtract_values(x,y):
# z = x - y
# return z
import my_functions
---------------------------------------------------------------------------
ModuleNotFoundError Traceback (most recent call last)
/tmp/ipykernel_2136/1130199131.py in <module>
----> 1 import my_functions
ModuleNotFoundError: No module named 'my_functions'
result = my_functions.subtract_values(10,5)
print(result)
